Bacteria are all around us — and that’s okay
Although these microbes remain poorly understood, they could prove key to protecting life across the planet
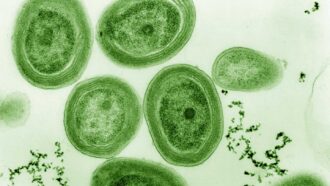
From the great outdoors to our internal organs, the world is awash in unseen bacteria (some seen growing on plate here). Most people assume (unfairly) that these germs are all dangerous. Biologists know better. Studying these poorly understood microbes could better reveal how they function as the "invisible backbone of life."
jarun011/iStockphoto
Victoria Orphan has loved the ocean for as long as she can remember. She used to snorkel in the Pacific Ocean near her family’s home in San Diego, Calif. She’d grab her mask and snorkel tube to visit the hidden world of plants and animals beneath the ocean’s surface. Orphan went to college at the University of California, Santa Barbara in the early 1990s. There she discovered something that changed the way she thought about the oceans — and life on Earth.
Another student showed her a small vial of seawater. Orphan didn’t think it looked all that interesting. It was just plain old water. Then the other student added a fluorescent chemical to the water and shined ultraviolet light on it. The tube lit up as millions of tiny bacteria began to glow. Just moments earlier, the microbes had been invisible. “These tiny organisms were all over the place,” says Orphan, “and yet we couldn’t see them. We knew almost nothing about them.”
She now spends her days exploring this hidden single-celled world. As a geobiologist at Caltech in Pasadena, Calif., she studies how bacteria and other microscopic life shape the deep sea.

Bacteria play central roles in many ecosystems. These include the oceans, soil and atmosphere. They’re also a big part of the global food web. Bacteria make it possible for all other life on Earth to exist. That’s why scientists say these single-celled organisms are the invisible backbone of all life — at least on Earth.
Yet there’s plenty we don’t know about them. Scientists think they’ve identified fewer than one percent of all bacterial species. That’s been driving Orphan and others to explore the mysteries of their one-celled world. They suspect bacteria will prove key to understanding — and protecting — Earth’s most important natural resources.
The methane eaters
Some bacteria eat really weird things. Scientists have found bacteria that eat rocks, sewage — even nuclear waste. Orphan studies a type of bacteria that live on the sea floor and gobble up methane.
Methane is a greenhouse gas. Like carbon dioxide and some other greenhouse gases, it enters the air when people burn oil, gas and coal. There are also natural sources of methane, such as natural gas, rice production and cow manure. Greenhouse gases trap heat in the atmosphere. An excess of these gases in Earth’s atmosphere has been warming the global climate.
Methane can seep out of the Earth on the sea floor. Some scientists say that even more methane would escape into the atmosphere if it wasn’t for marine bacteria. Certain of those bacteria dine on methane. That allows the oceans to trap a huge amount of the gas. “These microorganisms are the gatekeepers. They prevent ocean methane from getting into the atmosphere where it can change greenhouse-gas levels,” Orphan explains.
Finding single-celled organisms on the vast sea floor can be a challenge. Through the window of a submarine, she looks for clusters of clams and giant tube worms. These organisms signal that invisible marine bacteria live there, too. Wherever those methane-eaters live, they create new molecules as they dine. Other organisms use those new molecules as food. An entire food web springs up on the ocean floor.
Orphan and her team have found methane-eating bacteria along cracks on the sea floor, where this gas is seeping out. These cracks often happen where two tectonic plates bump into each other.
Some bacteria, they learned, can eat methane only by partnering with other single-celled organisms called archaea (Ar-KEE-uh). That important detail could help scientists better predict how much methane is escaping into the air, says Orphan.
In the trenches
Methane eaters aren’t the only deep-sea bacteria to interest scientists. “The deep sea is home to some pretty cool microbes,” says Jennifer Biddle. She’s a marine microbiologist at the University of Delaware in Newark. Biddle studies bacteria that live in deep ocean trenches.
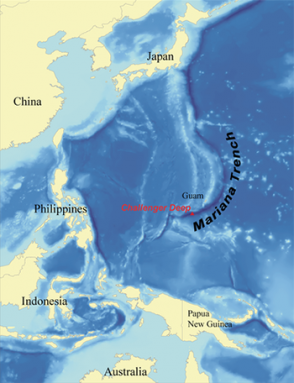
These underwater canyons are some of the least-studied places on Earth. They are incredibly hard to reach. Challenger Deep wins the record for the deepest-known spot on the planet. At the bottom of the Mariana Trench, in the western Pacific, Challenger Deep sits some 11 kilometers (more than 7 miles) below the ocean surface. If Mount Everest, the world’s tallest mountain, sat in the Mariana Trench, its peak would still be more than a mile beneath the waves.
The Mariana Trench is one of the toughest places for life to survive. Zero sunlight reaches it. Its temperatures are frigid. Large animals, such as whales or fish, can’t visit because the intense pressures there would crush them. Little surprise, then, that most of the locals are microscopic. They have adapted its extreme conditions.
Biddle and other scientists teamed up with deep-ocean explorers to send a submarine to Challenger Deep. James Cameron piloted the vessel. (He’s the movie director famous for Avatar and Titanic.) Cameron visited the bottom of Challenger Deep in March 2012 while making a documentary called Deepsea Challenge 3D. But the sub’s trek wasn’t just to get mesmerizing video for the Big Screen. The vessel also brought back sediment from the bottom of the trench.
Biddle and the other scientists screened that sediment for DNA. They were scouting for genes of familiar bacteria. They turned up evidence of some known as Parcubacteria.
Scientists didn’t even know this big group of bacteria existed until 2011. Back then, they found some in groundwater and dirt from a few places on land. But Biddle’s group now showed it also survives in one of the most inaccessible depths of the ocean.
Here, on the trench floor, the microbes were breathing nitrogen, not oxygen (as they did on land). And that makes sense. They had adapted to nitrogen since their home had little access to oxygen.
The more places we find such little-known bacteria, says Biddle, the more we can learn about what they do for their ecosystems.
Story continues below video.
From bread to biofuels
Even the bacteria in our kitchens and compost heaps interest scientists.
Sourdough bread gets its unique tart flavor when a mix of bacteria munches on the sugars in bread flour. Those bacteria make carbon dioxide, acids and other flavorful compounds. But to function, sourdough bacteria need their friends. Isolate just one bacterial species from the mix and the chemical reaction won’t happen. No sourdough.
Microbiologist Steve Singer lives near San Francisco, a California city famous for sourdough bread. He works for the Department of Energy at Lawrence Berkeley National Laboratory. And he suspected he could use the lessons of sourdough to make better biofuels. These plant-based fuels can power cars or trucks. They are considered “green,” meaning more Earth-friendly, than fossil fuels.

To make biofuels, scientists must break down plants into sugars. These sugars can then be turned into fuels such as ethanol (a type of alcohol). The chemical reactions that break down the plants require help from enzymes. These are molecules that jump-start or speed up chemical reactions.
The enzymes currently used to make biofuels are expensive. They also don’t work well, Singer says. That’s why researchers all over the world are searching for enzymes that might lower the cost and speed the production of biofuels.
He turned his search for them to the compost pile. There, bacterial communities were hard at work breaking down rotting fruits and veggies.
Singer took a small sample of the compost back to his lab. There, he let bacteria from the compost grow in a beaker. Later, he collected enzymes that these bacteria made and tested them on other plant bits. It worked: The enzymes broke down the plants into sugars.
Just as the sourdough bacteria need their friends to function, Singer discovered that these microbes produced the useful enzymes only when they were part of robust communities of different compost bacteria. Singer is now scaling up his project. His team is growing bacteria in huge vats called bioreactors. After he makes lots of the new enzymes, he can test whether they work better than existing ones for converting plant wastes into fuels.
“Taking something from the environment and trying to figure out how it works is one of the best parts of being a microbiologist,” Singer says.
Meta microbes
Singer is studying his new enzymes without knowing which bacteria are making them. This isn’t all that unusual. Bacteria are invisible to the unaided eye. Even with a microscope, telling two species apart can be hard. They don’t look as different as might two species of birds or flowers.
Scientists needed a different way to tell bacteria apart and know when they’ve stumbled upon new ones. Key to this sleuthing: DNA.
All organisms shed a little DNA throughout their environment. “It’s like a fingerprint. Each is unique,” explains Kelly Ramirez. She studies bacteria at the Netherlands Institute of Ecology in Wageningen.
Swab your kitchen counter and you might find human DNA (from you and your parents). There might be some plant DNA (from the veggies you just cut up) and from a fungus or two. There might even be some dog or cat DNA if you have a pet. You’ll also get a bunch of bacterial DNA because, well, bacteria a everywhere!
All of the cast-off genetic bits are known as environmental DNA, or eDNA.
Scientists can use these genetic fingerprints to discover new bacteria, notes Ramirez. They just need to bring any little bit of dirt or seawater or compost to a lab and check out what’s in it.
The sum of all the genetic material in an environment is called the metagenome (MET-uh-GEE-noam). Think of it as a DNA soup. All the molecules used to build the genes of different organisms are jumbled together.
Scientists use computers to untangle the mess.
Like a sieve, computer programs filter the soup. They look for familiar patterns known as genetic sequences. They form an organism’s DNA fingerprint. If scientists find a fingerprint they don’t recognize, it may be because it’s from some new species.
Scientists can compare these patterns to the fingerprints of familiar bacteria to see where the new bacteria fall within the tree of life. “We can now discover new microbes without ever seeing them,” explains Biddle at the University of Delaware.
The bacterial limb of the tree of life is sprouting new shoots and branches faster than at any time in history. Thirty years ago, all known single-celled organisms on the planet fit into a dozen major groups. Now there are about 120 known groups, or phyla (FY-lah). And the number of named bacteria in each group grows daily.
Little life, big data
What do you get when you add up the DNA sequences of millions of new bacteria? Lots and lots of data.
You can think about the planet as a machine, and all the ecosystems on Earth as the machine’s parts, says Jack Gilbert. All these data on bacterial DNA are key to “understanding the parts that make up the machine and how they all work together,” he says. Gilbert is a microbiologist at Argonne National Laboratory near Chicago, Ill.
His team is trying to organize those data into a virtual catalog of all the bacteria on Earth. It’s called the Earth Microbiome Project. More than 1,000 scientists around the world are helping collect samples. They’re looking in a host of different environments, then testing them for bacterial DNA.
So far the researchers have collected 100,000 samples. They’ve catalogued bacteria from the deepest ocean. They’ve found bacteria on the International Space Station, some 350 kilometers (220 miles) above Earth. They’ve discovered bacteria in exotic locations like the Amazon rainforest and ordinary places like public toilets.
Discovering which bacteria lurk there — and why — is the first step to understanding how different ecosystems drive the vast machine we think of as life on Earth. Learning about bacteria may help us answer questions about how our planet works, Gilbert says. Bacteria may explain why coral reefs in the ocean teem with life. Or they could explain why the soils of the North American prairie are so good for planting crops.
That’s why this search is so important, he says: “This is knowledge that can help us take better care of the planet.”