DNA in air can help ID unseen animals nearby
Looking for this wafting genetic material offers a new way to study animals

A new technique sucks DNA out of the air to successfully identify dozens of zoo animals, like this okapi.
Jurgen & Christine Sohns/Royalty free/Getty Images Plus
By Laura Allen
The okapi shot a dirty look at biologist Christina Islas Lynggaard. She was standing in his pen at Denmark’s Copenhagen Zoo using a noisy vacuum. Why? She hoped to collect bits of DNA he may have shed into the air.
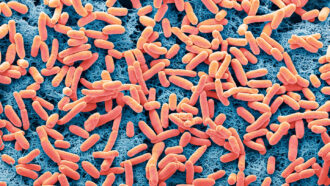
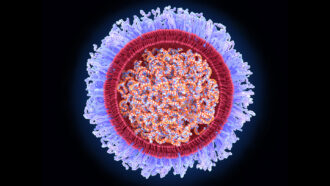
Living things shed traces of their DNA all the time. This genetic material comes out in saliva, fur, feces — even breath. It’s known as environmental DNA, or eDNA. For many years, scientists have used eDNA in water to identify what might be living in it.
Now, two research groups show that airborne eDNA can similarly identify nearby animals — even ones that can’t be seen.
Both groups shared their findings separately on January 6 in Current Biology.

Other researchers had previously tried sampling for bird eDNA in air. But until now, monitoring land animals has relied mostly on observation. To know what animals were in an area, researchers had to see critters with their eyes or snap photos of them with an automated camera (also known as a camera trap). This is time-consuming. It also can be especially hard when animals are shy or live in burrows.
DNA floating in air can help scientists study rare animals, including cryptic species — those who tend to hide well. Such eDNA also may make it easier and faster to monitor known communities of animals in the wild.
Educators and Parents, Sign Up for The Cheat Sheet
Weekly updates to help you use Science News Explores in the learning environment
Thank you for signing up!
There was a problem signing you up.
Zoos proved a good test site
Zoos contain many animals not found nearby in the wild. So the researchers decided to test their new strategy by looking for DNA from exotic species — perhaps a tiger — that clearly would not be roaming adjacent urban neighborhoods. A British group sampled air for such exotic beasts at Hamerton Zoo near Cambridge, England. Lynggaard’s group worked at the zoo in Denmark’s capital.
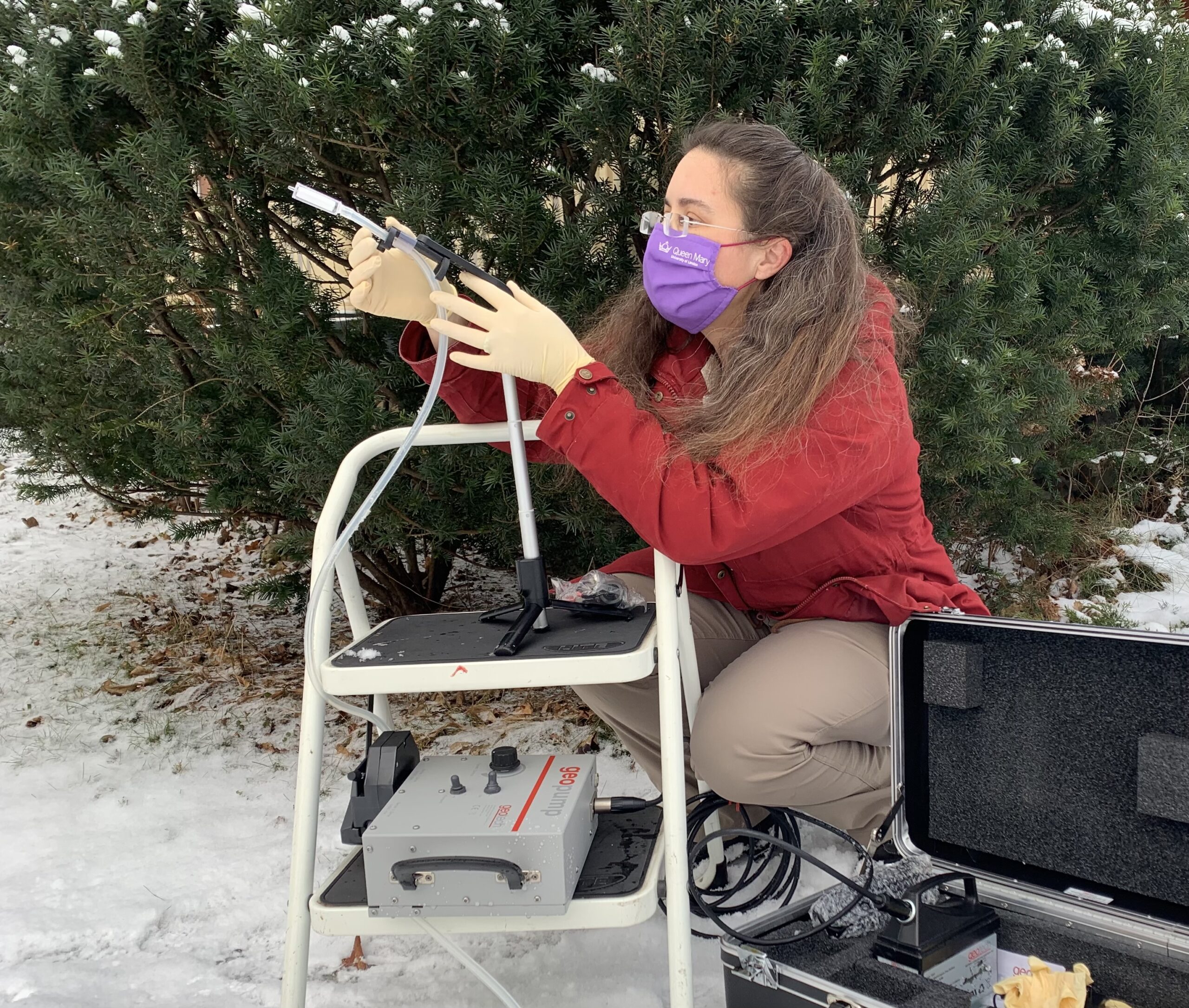
The group in Denmark used vacuums to collect DNA from the air. They tried a commercial vacuum, but it was loud and used lots of power. They ended up relying on a homemade model. The device was as small as a golf ball and quiet as a mouse, built from a little blower fan, a 3-D printed case and a piece of filter. The researchers sampled air in two enclosed areas: the okapi stable and the rainforest house containing frogs, sloths, armadillos and many birds. They also sampled at an outdoor site.

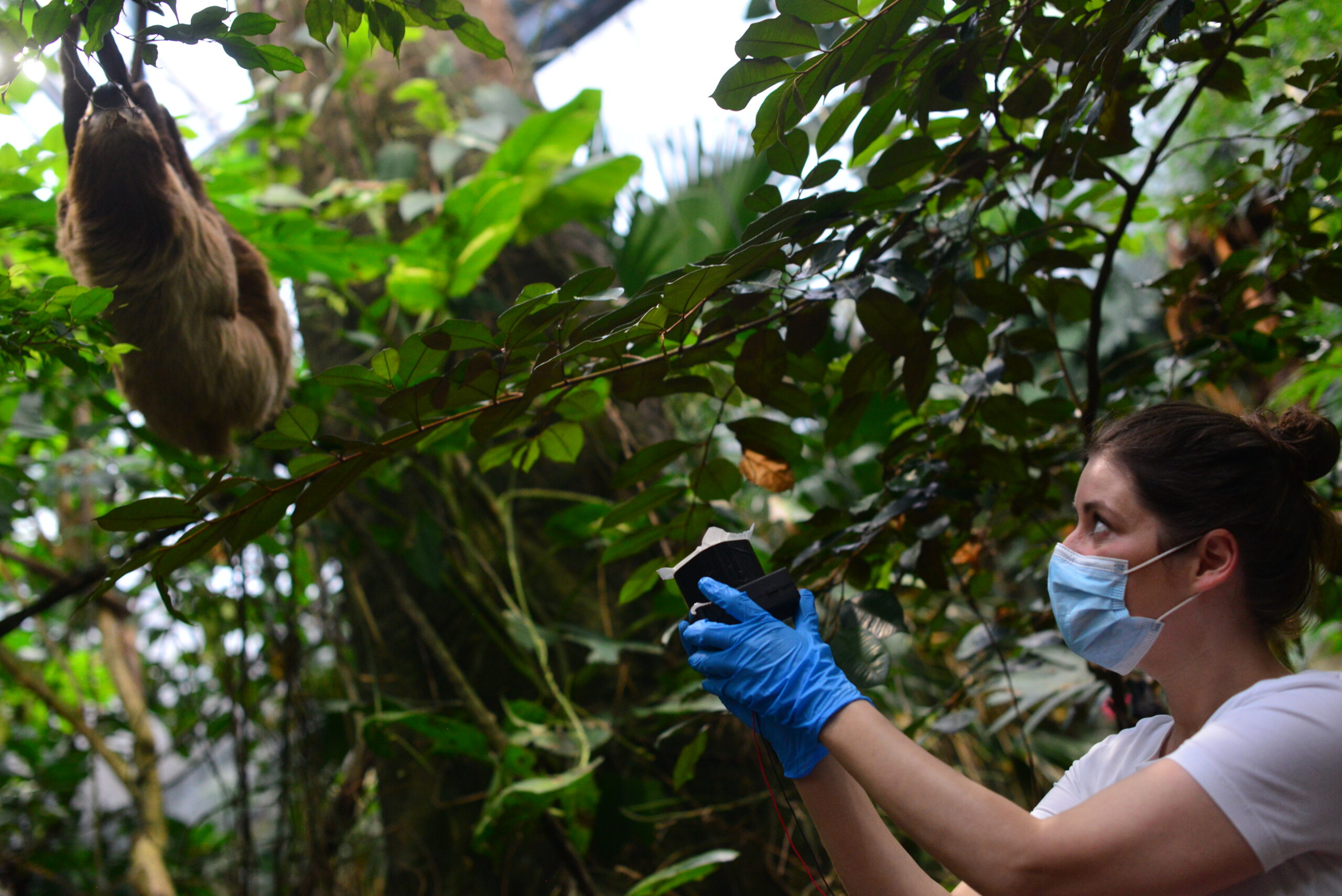
The team in England was led by Elizabeth Clare. She is a biologist who now works at York University in Toronto, Canada. Her team used a small pump to suck air through a very tiny filter.
Clare’s team did its work during the COVID-19 lockdown. The zoo had been closed for weeks. Without any visitors, most animals seemed excited to see the scientists.
“They’d follow us around their enclosures,” Clare says. A few “stole our equipment several times.” In fact, she had a tug-o-war with a binturong — a catlike Asian carnivore — over some electrical cords that it grabbed through its cage. In all, her team sampled at 20 sites around the zoo.
Both the pump and vacuum devices pulled incoming air through a filter, which collected any floating DNA.
Both teams took their filters back to the lab. All had DNA from bacteria and viruses together with DNA shed by plants and animals. The scientists selected just the DNA from vertebrate species, then made many copies of it. They did this using a process called polymerase (Puh-LIM-er-ace) chain reaction, or PCR. (It’s also what forensic scientists use when studying DNA found at a crime scene.) Next, they analyzed the copied DNA and compared its sequences against a database from known species. This told them which animals’ genetic material had been wafting around the collection site.
What they found
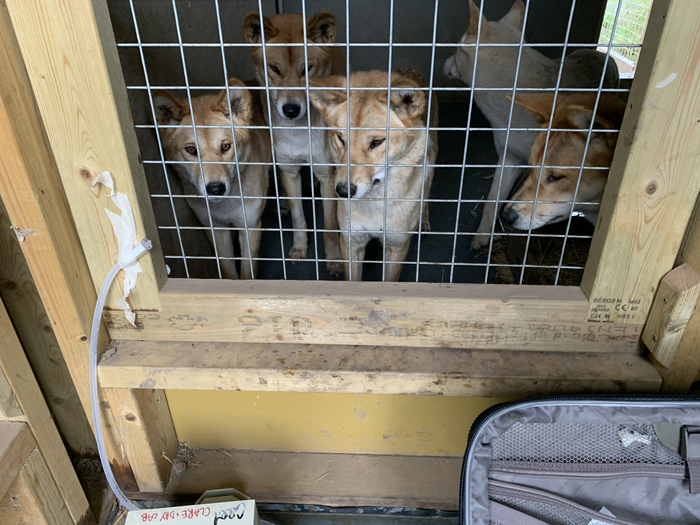
Clare’s team turned up DNA from vertebrates in 64 of its 72 samples. Most filters had DNA from multiple species. In all, the researchers detected 25 different types of animals, including a sloth, tiger, meerkat and dingo. These lived in the zoo. But not all of the vertebrate DNA came from exotic animals. Chicken DNA in the binturong enclosure, for instance, came from animals that had been fed to the zoo inhabitants. DNA from nearby wildlife also showed up. Most exciting to the U.K. team was DNA from the Eurasian hedgehog. It’s an endangered species in Britain.

In general, Clare’s group detected more DNA in samples collected closest to an animal. But the “sampled” animal didn’t have to be too close. The researchers detected meerkat DNA, for instance, from the dingo enclosure 245 meters (800 feet) away. This shows that DNA travels in the air and can be detected quite a distance from its source.
Similarly, air that Lynggaard vacuumed from the okapi pen contained DNA from 23 vertebrate species. These were zoo animals from other enclosures, wild animals and feeder fish. Throughout its zoo sampling, the team detected 30 mammals, 13 birds, four fish and one reptile species. They even detected tiny guppies from the pond inside the rainforest enclosure. “I was surprised we could detect them from the air,” she says.
The results are exciting other scientists. Masayuki Ushio is an ecologist at Kyoto University in Japan. He has used eDNA from water in his work. He now thinks air collection is “a promising approach” to monitor the diversity of species on land. He suspects it “will be an important contribution to conservation ecology.”
There is a lot to learn about DNA in the air. “I would expect that some species are more or less detectable,” says Lynggaard. “Maybe ones that groom all the time are easier to detect because they release more skin cells.”
“If I detect a tiger in a forest, I don’t know when it was there,” says Clare. “Yesterday? A week ago? A month ago? We don’t know how long the signal lasts.” And does it matter if it is warm or cold? Sunny or rainy? Now, she says, “lots of different teams of researchers are trying to answer these questions all over the world.”